 |
Fine DA, Rozenblatt-Rosen O, Padi M, Korkhin A, James RL, Adelmant G, Yoon R, Guo L, Berrios C, Zhang Y, Calderwood MA, Velmurgan S, Cheng J, Marto JA, Hill DE, Cusick ME, Vidal M, Florens L, Washburn MP, Litovchick L, DeCaprio JA..
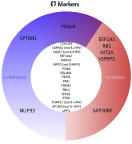
Identification of FAM111A as an SV40 host range restriction factor.
PLoS Pathog 2012; 8:e1002949.
|

|
 |
Neveu G, Cassonnet P, Vidalain PO, Rolloy C, Mendoza J, Jones L, Tangy F, Muller M, Demeret C, Tafforeau L, Lotteau V, Rabourdin-Combe C, Travé G, Dricot A, Hill DE, Vidal M, Favre M, Jacob Y.
Comparative analysis of virus-host interactomes with a mammalian high-throughput protein complementation assay based on Gaussia princeps luciferase.
Methods 2012; 58:349-59.
|

|
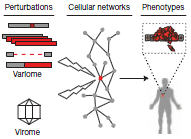 |
Rozenblatt-Rosen O, Deo RC, Padi M, Adelmant G, Calderwood MA, Rolland T, Grace M, Dricot A, Askenazi M, Tavares M, Pevzner SJ, Abderazzaq F, Byrdsong D, Carvunis AR, Chen AA, Cheng J, Correll M, Duarte M, Fan C, Feltkamp MC, Ficaro SB, Franchi R, Garg BK, Gulbahce N, Hao T, Holthaus AM, James R, Korkhin A, Litovchick L, Mar JC, Pak TR, Rabello S, Rubio R, Shen Y, Singh S, Spangle JM, Taşan M, Wanamaker S, Webber JT, Roecklein-Canfield J, Johannsen E, Barabási AL, Beroukhim R, Kieff E, Cusick ME, Hill DE, Münger K, Marto JA, Quackenbush J, Roth FP, DeCaprio JA, Vidal M.
Interpreting cancer genomes using systematic host network perturbations by tumour virus proteins.
Nature 2012; 487:491–5.
|

|
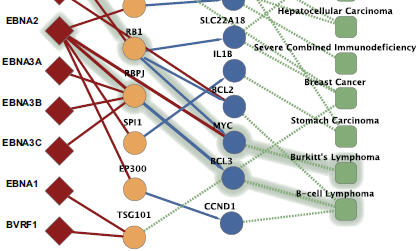 |
Gulbahce N, Yan H, Dricot A, Padi M, Byrdsong D, Franchi R, Lee DS, Rozenblatt-Rosen O, Mar JC, Calderwood MA, Baldwin A, Zhao B, Santhanam B, Braun P, Huh KW, Hellner K, Grace M, Chen A, Rubio R, Marto JA, Christakis NA, Kieff E, Roth FP, Roecklein-Canfield J, DeCaprio JA, Cusick ME, Quackenbush J, Hill DE, Münger K, Vidal M, Barabási AL.
Viral perturbations of host networks reflect disease etiology.
PLoS Comput Biol 2012; 8:e1002531.
|

|
 |
Heilmann AM, Calderwood MA, Portal D, Lu Y, Johannsen E.
Genome-wide analysis of Epstein-Barr virus rta DNA binding.
J Virol 2012; 86:5151-64.
|

|
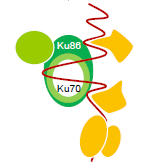 |
Adelmant G, Calkins AS, Garg BK, Card JD, Askenazi M, Miron A, Sobhian B, Zhang Y, Nakatani Y, Silver PA, Iglehart JD, Marto JA, Lazaro JB.
DNA ends alter the molecular composition and localization of Ku multi-component complexes.
Mol Cell Proteomics 2012; 11:411-21.
|

|
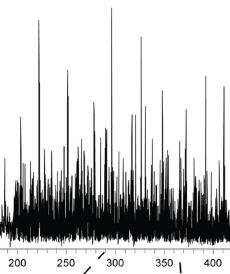 |
Zhou F, Lu Y, Ficarro SB, Webber JT, Marto JA.
Nanoflow low pressure high peak capacity single dimension LC-MS/MS platform for high-throughput, in-depth analysis of mammalian proteomes.
Anal Chem 2012; 84:5133-9.
|

|
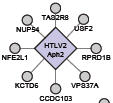 |
Simonis N, Rual JF, Lemmens I, Boxus M, Hirozane-Kishikawa T, Gatot JS, Dricot A, Hao T, Vertommen D, Legros S, Daakour S, Klitgord N, Martin M, Willaert JF, Dequiedt F, Navratil V, Cusick ME, Burny A, Lint CV, Hill DE, Tavernier J, Kettmann R, Vidal M, Twizere JC.
Host-pathogen interactome mapping for HTLV-1 and -2 retroviruses.
Retrovirology 2012; 9:26.
|

|
 |
Cardoza JD, Parikh JR, Ficarro SB, Marto JA.
Mass spectrometry-based proteomics: qualitative identification to activity-based protein profiling.
Wiley Interdiscip Rev Syst Biol Med 2012; 4:141-62.
|

|
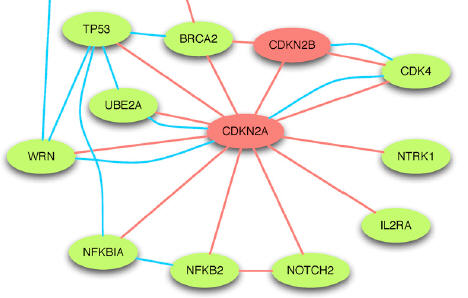 |
Taşan M, Drabkin HJ, Beaver JE, Chua HN, Dunham J, Tian W, Blake JA, Roth FP.
A resource of quantitative functional annotation for Homo sapiens genes.
Genes Genomes Genet 2012; 2:223-33.
|

|
 |
Culhane AC, Schröder MS, Sultana R, Picard SC, Martinelli EN, Kelly C, Haibe-Kains B, Kapushesky M, St Pierre AA, Flahive W, Picard KC, Gusenleitner D, Papenhausen G, O’Connor N, Correll M, Quackenbush J.
GeneSigDB: a manually curated database and resource for analysis of gene expression signatures.
Nucleic Acids Res 2012; 40:D1060-6.
|

|
 |
Schröder MS, Culhane AC, Quackenbush J, Haibe-Kains B.
survcomp: an R/Bioconductor package for performance assessment and comparison of survival models.
Bioinformatics 2011; 27:3206-8. |

|
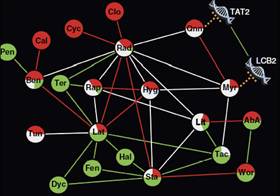 |
Cokol M, Chua HN, Taşan M, Mutlu B, Weinstein ZB, Suzuki Y, Nergiz ME, Costanzo M, Baryshnikova A, Giaever G, Nislow C, Myers CL, Andrews BJ, Boone C, Roth FP.
Systematic exploration of synergistic drug pairs.
Mol Syst Biol 2011; 7:544. |

|
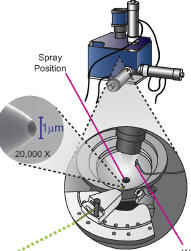 |
Ficarro SB, Zhang Y, Carrasco-Alfonso MJ, Garg B, Adelmant G, Webber JT, Luckey CJ, Marto JA.
Online nanoflow multidimensional fractionation for high efficiency phosphopeptide analysis.
Mol Cell Proteomics 2011; 10:O111.011064. |

|
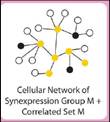 |
Mar JC, Matigian NA, Quackenbush J, Wells CA.
attract: a method for identifying core pathways that define cellular phenotypes.
PLoS ONE 2011; 6:e25445. |

|
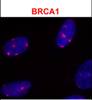 |
Pathania S, Nguyen J, Hill SJ, Scully R, Adelmant GO, Marto JA, Feunteun J, Livingston DM.
BRCA1 is required for postreplication repair after UV-induced DNA damage.
Mol Cell 2011; 44:235-51. |

|
 |
Zhou F, Sikorski TW, Ficarro SB, Webber JT, Marto JA..
Online nanoflow reversed phase-strong anion exchange-reversed phase liquid chromatography-tandem mass spectometry platform for effecient and in-depth proteome sequence analysis of complex organism.
Anal Chem 2011; 83:6996-7005. |

|
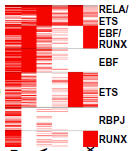 |
Zhao B, Zou J, Wang H, Johannsen E, Peng CW, Quackenbush J, Mar JC, Morton CC, Freedman ML, Blacklow SC, Aster JC, Bernstein BE, Kieff E.
Epstein-Barr virus exploits intrinsic B-lymphocyte transcription programs to achieve immortal cell growth.
Proc Natl Acad Sci USA 2011; 108:14902-7. |

|
 |
Mar JC, Matigian NA, Mackay-Sim A, Mellick GD, Sue CM, Silburn PA, McGrath JJ, Quackenbush J, Wells CA.
Variance of gene expression identifies altered network constraints in neurological disease.
PLoS Genet 2011; 7:e1002207. |

|
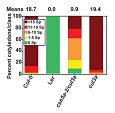 |
Mukhtar MS, Carvunis AR, Dreze M, Epple P, Steinbrenner J, Moore J, Taşan M, Galli M, Hao T, Nishimura MT, Pevzner SJ, Donovan SE, Ghamsari L, Santhanam B, Romero V, Poulin MM, Gebreab F, Gutierrez BJ, Tam S, Monachello D, Boxem M, Harbort CJ, McDonald N, Gai L, Chen H, He Y, Vandenhaute J, Roth FP, Hill DE, Ecker JR, Vidal M, Beynon J, Braun P, Dangl JL.
Independently evolved virulence effectors converge onto hubs in a plant immune system network.
Science 2011; 333:596-601. |

|
 |
Gewurz BE, Mar JC, Padi M, Zhao B, Shinners NP, Takasaki K, Bedoya E, Zou JY, Cahir-McFarland E, Quackenbush J, Kieff E.
Canonical NF-κB activation is essential for Epstein-Barr virus latent membrane protein 1 TES2/CTAR2 gene regulation.
J Virol 2011; 85:6764-73. |

|
 |
Portal D, Zhao B, Calderwood MA, Sommermann T, Johannsen E, Kieff E.
EBV nuclear antigen EBNALP dismisses transcription repressors NCoR and RBPJ from enhancers and EBNA2 increases NCoR-deficient RBPJ DNA binding.
Proc Natl Acad Sci USA 2011; 108:7808-13. |

|
 |
Askenazi M, Webber JT, Marto JA.
mzServer: web-based programmatic access for mass spectrometry data analysis.
Mol Cell Proteomics 2011; 10:M10.003988. |

|
 |
Webber JT, Askenazi M, Marto JA.
mzResults: an interactive viewer for interrogation and distribution of proteomics results.
Mol Cell Proteomics 2011; 10:M110.003970. |

|
 |
Hellner K, Münger K.
Human papillomavirus as therapeutic targets in human cancer.
J Clin Oncol 2011; 29:1785-94. |

|
 |
Mar JC, Wells CA, Quackenbush J.
Defining an informativeness metric for clustering gene expression data.
Bioinformatics 2011; 27:1094-100. |

|
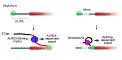 |
Cenik C, Chua HN, Zhang H, Tarnawsky SP, Akef A, Derti A, Taşan M, Moore MJ, Palazzo AF, Roth FP.
Genome analysis reveals interplay between 5’UTR introns and nuclear mRNA export for secretory and mitochondrial genes.
PLoS Genet 2011; 7:e1001366. |

|
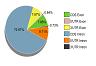 |
Sephton CF, Cenik C, Kucukural A, Dammer EB, Cenik B, Han Y, Dewey CM, Roth FP, Herz J, Peng J, Moore MJ, Yu G.
Identification of neuronal RNA targets of TDP-43-containing ribonucleoprotein complexes.
J Biol Chem 2011; 286:1204-15. |

|
 |
Zhao B, Mar JC, Maruo S, Lee S, Gewurz BE, Johannsen E, Holton K, Rubio R, Takada K, Quackenbush J, Kieff E.
Epstein-Barr virus nuclear antigen 3C regulated genes in lymphoblastoid cell lines.
Proc Natl Acad Sci USA 2011; 108:337-42. |

|
 |
Dreze M, Monachello D, Lurin C, Cusick ME, Hill DE, Vidal M, Braun P.
High-quality binary interactome mapping.
Methods Enzymol 2011; 470:281-315. |

|
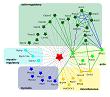 |
Meixner A, Boldt K, Van Troys M, Askenazi M, Gloeckner CJ, Bauer M, Marto JA, Ampe C, Kinkl N, Ueffing M.
A QUICK screen for Lrrk2 interaction partners–leucine-rich repeat kinase 2 is involved in actin cytoskeleton dynamics.
Mol Cell Proteomics 2011; 10:M110.001172. |

|
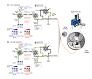 |
Zhou F, Cardoza JD, Ficarro SB, Adelmant GO, Lazaro JB, Marto JA.
Online nanoflow RP-RP-MS reveals dynamics of multicomponent Ku complex in response to DNA damage.
J Proteome Res 2010; 9:6242-55. |

|
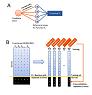 |
Yan H, Venkatesan K, Beaver JE, Klitgord N, Yildirim MA, Hao T, Hill DE, Cusick ME, Perrimon N, Roth FP, Vidal M.
A genome-wide gene function prediction resource for Drosophila melanogaster.
PLoS ONE 2010; 5:e12139. |

|
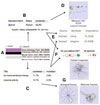 |
Beaver JE, Tasan M, Gibbons FD, Tian W, Hughes TR, Roth FP.
FuncBase: a resource for quantitative gene function annotation.
Bioinformatics 2010; 26:1806-7. |

|
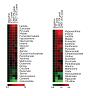 |
Lewis GD, Farrell L, Wood MJ, Martinovic M, Arany Z, Rowe GC, Souza A, Cheng S, McCabe EL, Yang E, Shi X, Deo R, Roth FP, Asnani A, Rhee EP, Systrom DM, Semigran MJ, Vasan RS, Carr SA, Wang TJ, Sabatine MS, Clish CB, Gerszten RE.
Metabolic signatures of exercise in human plasma.
Sci Transl Med 2010; 2:33ra37. |

|
 |
Herrera M, Roberts DC, Gulbahce N.
Mapping the evolution of scientific fields.
PLoS ONE 2010; 5:e10355. |
 |
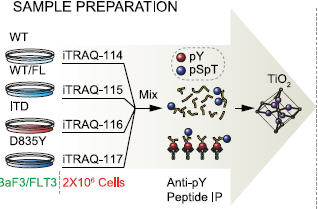 |
Zhang Y, Askenazi M, Jiang J, Luckey CJ, Griffin JD, Marto JA.
A robust error model for iTRAQ quantification reveals divergent signaling between oncogenic FLT3 mutants in acute myeloid leukemia.
Mol Cell Proteomics 2010; 9:780-90. |

|
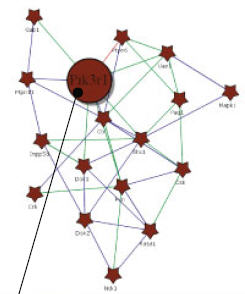 |
Askenazi M, Li S, Singh S, Marto JA.
Pathway Palette: a rich internet application for peptide-, protein- and network-oriented analysis of MS data.
Proteomics 2010; 10:1880-5. |

|
 |
Derti A, Cenik C, Kraft P, Roth FP.
Absence of evidence for MHC-dependent mate selection within HapMap populations.
PLoS Genet 2010; 6:e1000925. |
 |
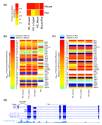 |
Cenik C, Derti A, Mellor JC, Berriz GF, Roth FP.
Genome-wide functional analysis of human 5′ untranslated region introns.
Genome Biol 2010; 11:R29. |

|
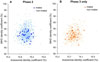 |
Deo RC, Hunter L, Lewis GD, Pare G, Vasan RS, Chasman D, Wang TJ, Gerszten RE, Roth FP.
Interpreting metabolomic profiles using unbiased pathway models.
PLoS Comput Biol 2010; 6:e1000692. |

|
 |
Culhane AC, Schwarzl T, Sultana R, Picard KC, Picard SC, Lu TH, Franklin KR, French SJ, Papenhausen G, Correll M, Quackenbush J.
GeneSigDB–a curated database of gene expression signatures.
Nucleic Acids Res 2010; 38:D716-25. |

|
 |
Shapira SD, Gat-Viks I, Shum BO, Dricot A, de Grace MM, Wu L, Gupta PB, Hao T, Silver SJ, Root DE, Hill DE, Regev A, Hacohen N.
A physical and regulatory map of host-influenza interactions reveals pathways in H1N1 infection.
Cell 2009; 139:1255-67. |

|
 |
Aryee MJ, Gutierrez-Pabello JA, Kramnik I, Maiti T, Quackenbush J.
An improved empirical bayes approach to estimating differential gene expression in microarray time-course data: BETR (Bayesian Estimation of Temporal Regulation).
BMC Bioinformatics 2009; 10:409. |

|
 |
Mar JC, Quackenbush J.
Decomposition of gene expression state space trajectories.
PLoS Comput Biol 2009; 5:e1000626. |

|
 |
Berriz GF, Beaver JE, Cenik C, Tasan M, Roth FP.
Next generation software for functional trend analysis.
Bioinformatics 2009; 25:3043-4. |

|
 |
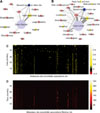
Parikh JR, Askenazi M, Ficarro SB, Cashorali T, Webber JT, Blank NC, Zhang Y, Marto JA.
multiplierz: an extensible API based desktop environment for proteomics data analysis.
BMC Bioinformatics 2009; 10:364. |

|
 |
Chen LL, Blumm N, Christakis NA, Barabasi AL, Deisboeck TS.
Cancer metastasis networks and the prediction of progression patterns.
Br J Cancer 2009; 101:749-58. |

|
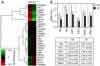 |
Hellner K, Mar J, Fang F, Quackenbush J, Munger K.
HPV16 E7 oncogene expression in normal human epithelial cells causes molecular changes indicative of an epithelial to mesenchymal transition.
Virology 2009; 391:57-63. |
 |
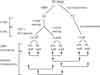 |
Deo RC, Roth FP.
Pathways of the heart.
Circ Cardiovasc Genet 2009; 2:303-5. |

|
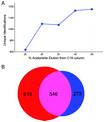 |
Ficarro SB, Adelmant G, Tomar MN, Zhang Y, Cheng VJ, Marto JA.
Magnetic bead processor for rapid evaluation and optimization of parameters for phosphopeptide enrichment.
Anal Chem 2009; 81:4566-75. |

|
 |
Mar JC, Kimura Y, Schroder K, Irvine KM, Hayashizaki Y, Suzuki H, Hume D, Quackenbush J.
Data-driven normalization strategies for high-throughput quantitative RT-PCR.
BMC Bioinformatics 2009; 10:110. |

|
 |
Park J, Lee DS, Christakis NA, Barabasi AL.
The impact of cellular networks on disease comorbidity.
Mol Syst Biol 2009; 5:262. |

|
 |
Hidalgo CA, Blumm N, Barabasi AL, Christakis NA.
A dynamic network approach for the study of human phenotypes.
PLoS Comput Biol 2009; 5:e1000353. |

|
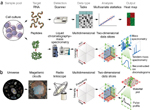 |
Askenazi M, Parikh JR, Marto JA.
mzAPI: a new strategy for efficiently sharing mass spectrometry data.
Nat Methods 2009; 6:240-1. |

|
 |
Adelmant G, Marto JA.
Protein complexes: the forest and the trees.
Expert Rev Proteomics 2009; 6:5-10. |

|
 |
Braun P, Tasan M, Dreze M, Barrios-Rodiles M, Lemmens I, Yu H, Sahalie JM, Murray RR, Roncari L, de Smet AS, Venkatesan K, Rual JF, Vandenhaute J, Cusick ME, Pawson T, Hill DE, Tavernier J, Wrana JL, Roth FP, Vidal M.
An experimentally derived confidence score for binary protein-protein interactions.
Nat Methods 2009; 6:91-7. |

|
 |
Venkatesan K, Rual JF, Vazquez A, Stelzl U, Lemmens I, Hirozane-Kishikawa T, Hao T, Zenkner M, Xin X, Goh KI, Yildirim MA, Simonis N, Heinzmann K, Gebreab F, Sahalie JM, Cevik S, Simon C, de Smet AS, Dann E, Smolyar A, Vinayagam A, Yu H, Szeto D, Borick H, Dricot A, Klitgord N, Murray 30. RR, Lin C, Lalowski M, Timm J, Rau K, Boone C, Braun P, Cusick ME, Roth FP, Hill DE, Tavernier J, Wanker EE, Barabasi AL, Vidal M.
An empirical framework for binary interactome mapping.
Nat Methods 2009; 6:83-90. |

|
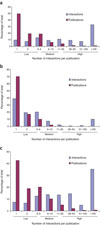 |
Cusick ME, Yu H, Smolyar A, Venkatesan K, Carvunis AR, Simonis N, Rual JF, Borick H, Braun P, Dreze M, Vandenhaute J, Galli M, Yazaki J, Hill DE, Ecker JR, Roth FP, Vidal M.
Literature-curated protein interaction datasets.
Nat Methods 2009; 6:39-46. |

|
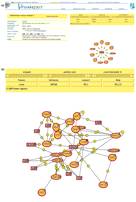 |
Chatr-aryamontri A, Ceol A, Peluso D, Nardozza A, Panni S, Sacco F, Tinti M, Smolyar A, Castagnoli L, Vidal M, Cusick ME, Cesareni G.
VirusMINT: a viral protein interaction database.
Nucleic Acids Res 2009; 37:D669-73. |

|
 |
Gulbahce N, Lehmann S.
The art of community detection.
Bioessays 2008; 30:934-8. |

|
 |
Lewis GD, Wei R, Liu E, Yang E, Shi X, Martinovic M, Farrell L, Asnani A, Cyrille M, Ramanathan A, Shaham O, Berriz G, Lowry PA, Palacios IF, Tasan M, Roth FP, Min J, Baumgartner C, Keshishian H, Addona T, Mootha VK, Rosenzweig A, Carr SA, Fifer MA, Sabatine MS, Gerszten RE.
Metabolite profiling of blood from individuals undergoing planned myocardial infarction reveals early markers of myocardial injury.
J Clin Invest 2008; 118:3503-12. |

|
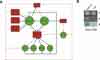 |
Martinez NJ, Ow MC, Barrasa MI, Hammell M, Sequerra R, Doucette-Stamm L, Roth FP, Ambros VR, Walhout AJ.
A C. elegans genome-scale microRNA network contains composite feedback motifs with high flux capacity.
Genes Dev 2008; 22:2535-49. |

|
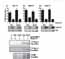 |
Calderwood MA, Holthaus AM, Johannsen E.
The Epstein-Barr virus LF2 protein inhibits viral replication.
J Virol 2008; 82:8509-19. |

|
 |
Lee DS, Park J, Kay KA, Christakis NA, Oltvai ZN, Barabasi AL.
The implications of human metabolic network topology for disease comorbidity.
Proc Natl Acad Sci USA 2008; 105:9880-5. |

|
 |
Pena-Castillo L, Tasan M, Myers CL, Lee H, Joshi T, Zhang C, Guan Y, Leone M, Pagnani A, Kim WK, Krumpelman C, Tian W, Obozinski G, Qi Y, Mostafavi S, Lin GN, Berriz GF, Gibbons FD, Lanckriet G, Qiu J, Grant C, Barutcuoglu Z, Hill DP, Warde-Farley D, Grouios C, Ray D, Blake JA, Deng M, Jordan MI, Noble WS, Morris Q, Klein-Seetharaman J, Bar-Joseph Z, Chen T, Sun F, Troyanskaya OG, Marcotte EM, Xu D, Hughes TR, Roth FP.
A critical assessment of Mus musculus gene function prediction using integrated genomic evidence.
Genome Biol 2008; 9 Suppl 1:S2. |

|
 |
Chittenden TW, Howe EA, Culhane AC, Sultana R, Taylor JM, Holmes C, Quackenbush J.
Functional classification analysis of somatically mutated genes in human breast and colorectal cancers.
Genomics 2008; 91:508-11. |

|
 |
Park J, Barabasi AL.
Distribution of node characteristics in complex networks.
Proc Natl Acad Sci USA 2007; 104:17916-20. |

|